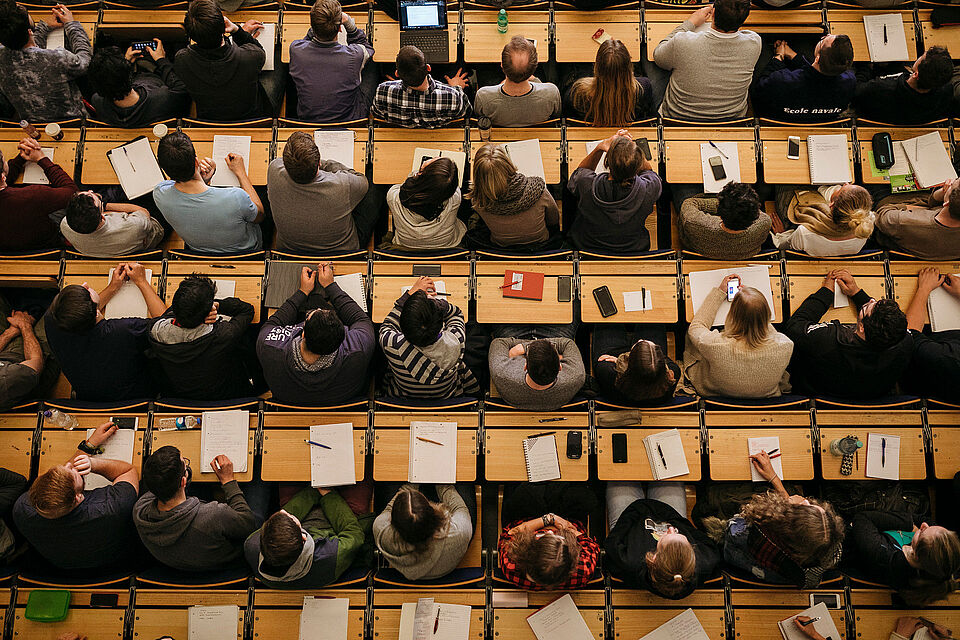
 © Alexander Rentsch
© Alexander Rentsch
Institut für Mathematik
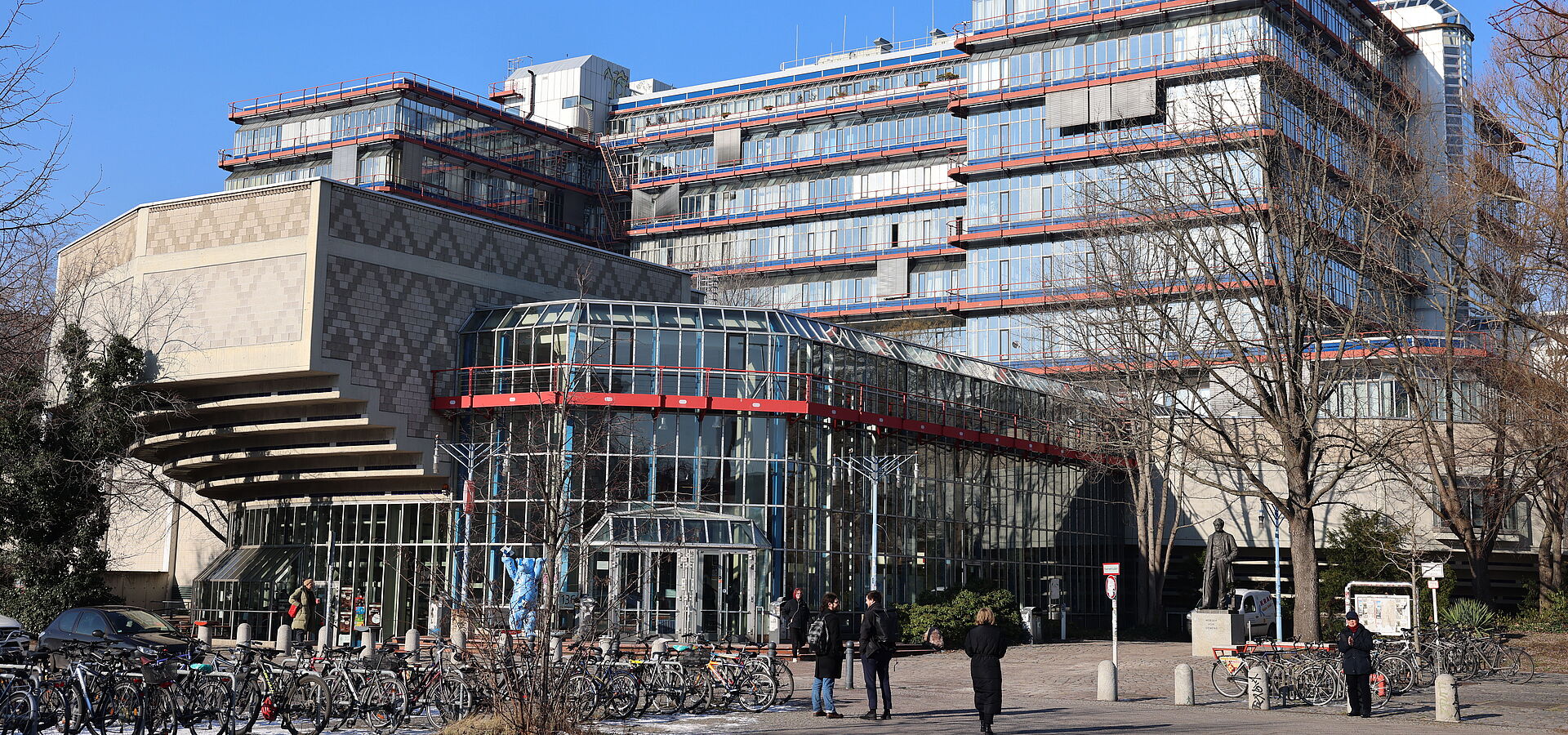

Willkommen
Das Institut für Mathematik (IfM) der Fakultät II – Mathematik und Naturwissenschaften – ist als Teil einer technischen Universität in Forschung und Lehre anwendungsnah orientiert. Es bietet die Studiengänge Mathematik, Wirtschaftsmathematik, Technomathematik (jeweils Bachelor und Master) und Scientific Computing (nur Master) sowie umfangreichen Mathematik-Service für Studierende anderer Studiengänge an.

 © Felix Noak
© Felix Noak
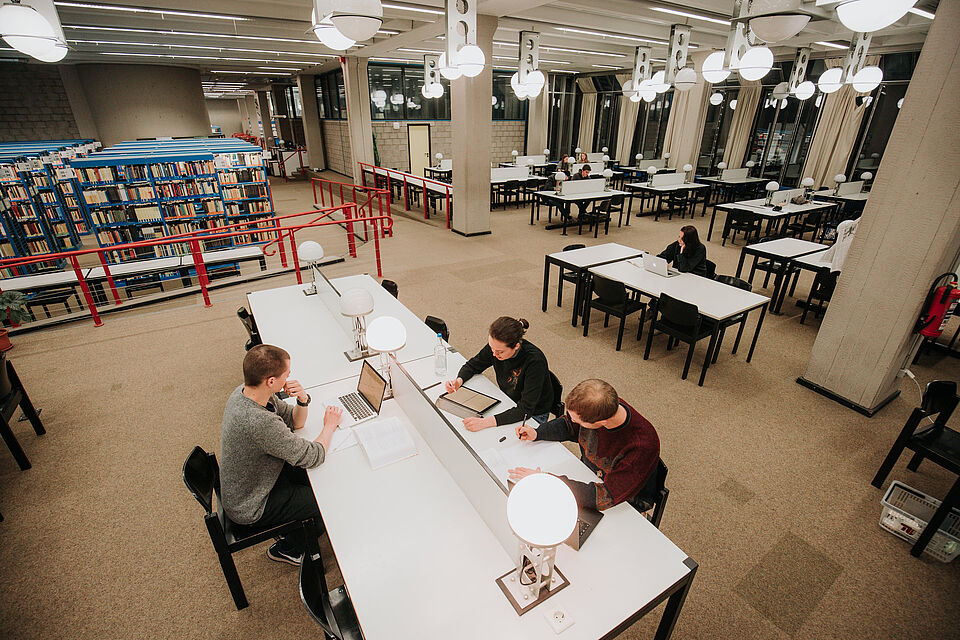


 © Oana Popa-Costea
© Oana Popa-Costea
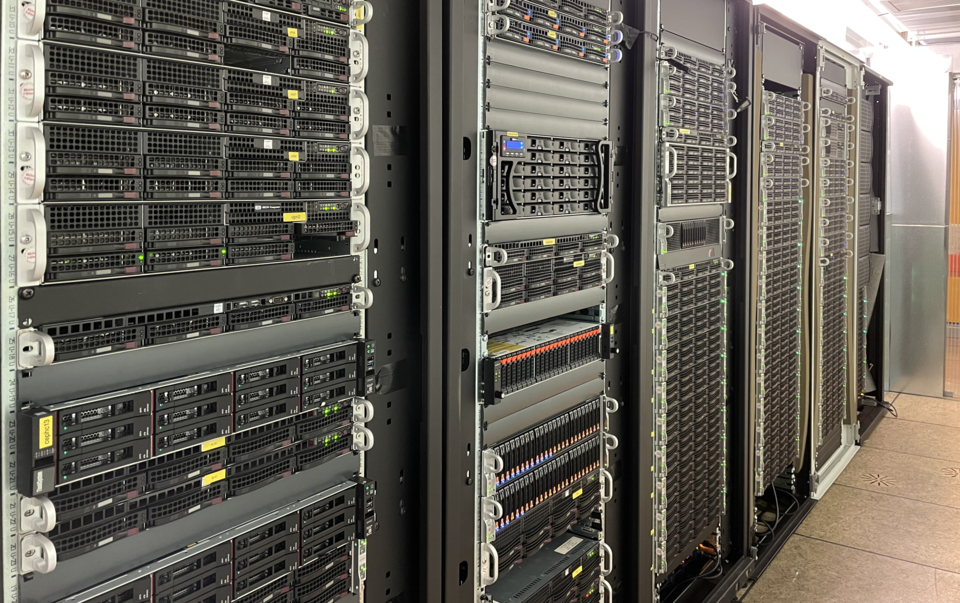 © A. Flügge
© A. Flügge